My previous works
1. Seasonal dominance of the CL500-11 lineage in the hypolimnion of Lake Biwa
The dominance of the CL500-11 lineage was discovered in the oxygenated hypolimnion of Lake Biwa. Spatiotemporal profiling of their abundance indicated their exclusive occurrence in the hypolimnion during the stratified period, concluding that CL500-11 is hypolimnion-specific bacterioplankton (Okazaki et al., 2013).
2. Vertical partitioning of bacterioplankton community in Lake Biwa
A spatiotemporal survey of bacterioplankton community composition in Lake Biwa revealed that the community was different at the phylum-level between the epilimnion and hypolimnion during the stratified period. The result allowed us to identify numbers of hypolimnion-specific lineages other than CL500-11 (Okazaki & Nakano 2016).
3. Comparative investigation of 10 Japanese deep freshwater lakes
Ubiquity and quantitative significance of the hypolimnion-specific lineages identified in Lake Biwa were confirmed by a comprehensive sampling in 10 deep Japanese freshwater lakes. The results established a first general overview of bacterioplankton lineages inhabiting the oxygenated hypolimnia of deep freshwater lakes (Okazaki et al., 2017).
4. Investigation of CL500-11 dominance in perialpine glacier lakes
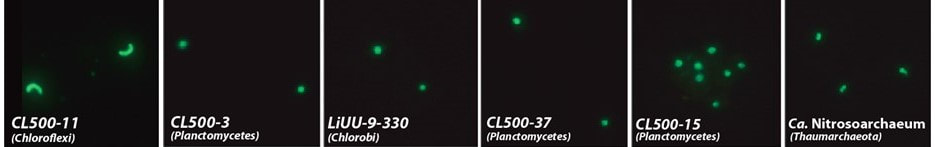
The dominance of the CL500-11 lineage was investigated by CARD-FISH in 7 deep perialpine glacier lakes. The result revealed that CL500-11 dominated in all investigated lakes. By summarizing the results together with published data, this study revealed their broad habitat spectrum and possible eco-physiological characteristics (Okazaki et al., 2018).
5. Shotgun metagenomics of bacteria and viruses in Lake Biwa
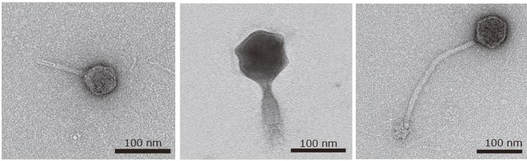
The genomic diversities of bacteria and viruses (phages) in Lake Biwa were revealed through spatiotemporal metagenomic analyses. The results characterized potential key virus-host interactions that are thought to play important roles in the ecosystem (Okazaki et al., 2019).
6. Resolving microdiversification of lake bacterioplankton using long-read amplicon sequencing
Intra-lineage diversities of predominant freshwater bacterioplankton lineages were revealed through a high-resolution, long-read amplicon sequencing analysis. The results showed that there is a clear intra-lineage geographic isolation among members inhabiting different lakes and revealed that the degree of microdiversification varies among lineages, possibly reflecting differences in their ecological strategies as a species (Okazaki et al., 2021).
7. Resolving genomic microdiversity of Lake Biwa bacterioplankton genomes
By targeting the bacterial assemblages in Lake Biwa and utilizing long-read sequencing and read-mapping analysis,, this study developed a metagenomic approach that can comprehensively detect slight variations (nucleotide and structural variants) in environmental bacterial genomes. Comparative analysis of the microdiversity among genomes and genes revealed factors behind microbial genomic diversification, namely, that diversification is driven primarily by resistance against viral infection and constrained by the population size. (Okazaki et al., 2022).